 ...the monthly, Open Access Publisher.
...the monthly, Open Access Publisher.
 ...the monthly, Open Access Publisher.
...the monthly, Open Access Publisher.
Research Article - (2025) Volume 12, Issue 1
Received: 23-Aug-2023, Manuscript No. IJPBG-23-111139; Editor assigned: 28-Aug-2023, Pre QC No. IJPBG-23-111139 (PQ); Reviewed: 12-Sep-2023, QC No. IJPBG-23-111139; Revised: 13-Jan-2025, Manuscript No. IJPBG-23-111139 (R); Published: 20-Jan-2025
Background and objective: The genetic structuring of populations could be explained by several parameters such as geographical distance and anthropogenic activities. The Niger date palm population is genetically structured into biogeographical distribution of subpopulations; however, it is important to find out the main causes of this structuring. Therefore, this study aims to assess whether this genetic structure is mainly attributable to the constraints related to the distribution of date palm in Niger.
Materials and methods: The matrix of genetic distances (Fst), the matrix of geographic distances, and the anthropic activities matrix (Euclidean distance), between subpopulations were estimated. Mantel's test was used to determine the relationships between these matrices. Indeed, the Mantel test allowed to evaluate the influence of geographic distance matrices and that of the implementation of cultural practices on the matrix of genetic differentiation between subpopulations. The observed variation was compared to the theoretical at 100,000 repetitions for better accuracy.
Results: This study showed a significant correlation between the genetic differentiation of subpopulations and the geographical distance between subpopulations (rm=0.280 and P-value=0.000) as well as with that of the implementation of cultural practices related to the date palm (rm=0.534 and P-value=0.000). The genetic variability between date palm subpopulations can be predicted according to the implementation of the cultural practices adopted by the producers and according to the geographical dispersion of the subpopulations. But the cause-and-effect analysis between genetic distances and geographic distances showed that this relationship is not directly causal, so it’s a spatial autocorrelation.
Conclusion: This study is a reference that provides knowledge on the genetic variation between Phoenix dactylifera subpopulations according to anthropic and geographical factors. This serves as information for the sustainable conservation of date palm wealth in Niger.
Date palm, Geographical isolation, Cultural practice, Genetic structure
Genetic variation between populations could be explained by processes like adaptation to local activities and environmental conditions [1]. The geographic distance limits genetic exchange between subpopulations, because the rate of migration is higher between the closest populations, lower between those that are further away [2]. The combination of local environmental conditions and genetic migration limited by the distance between subpopulations could act more on the local differentiation of allele frequencies which can considerably impact the genetic structure of populations. In the date palm areas of Niger, some palm groves are very far from each other and even though cultivation practices vary from one area to another and from one palm grove to another [3]. The palm groves of Niger are distributed in the Saharan and Sahelian zones and that there is a relationship between the geographical distribution and the cultivation practices [4]. Furthermore, the history of palm groves in Niger shows that the two subpopulations have a different origin. The palm groves of the Sahara are old (1570 years) and therefore more sophisticated cultivation practices have been developed. Conversely, those of the Sahel are recent (100 years) and few cultural practices are applied. The same information was reported by the study carried out in Chad by Oumar, et al. The implementation of certain cultural practices such as modes of multiplication and modes of reproduction, leads to the appearance of new phenotypes and therefore a genotypic variation which can be interesting but also has several disadvantages [5].
Genetic differentiation between subpopulations induced by geographic distance has been observed in many species [6], with a large literature devoted to identifying other environmental factors influencing diversity and genetic structuring of populations, but not yet on the date palm (Phoenix dactylifera). Thus, questions about the relationship between the genetic structure of populations and the environment are multiple. Can the understanding of the causes of genetic structure be improved by using geographical and cultural information’s? How to detect environmental and/or anthropic factors having a strong link with the genetic structure? Therefore, this study aims to:
Study area and data collection
Study area and vegetal material: The study area is Saharan and Sahelian zones, two main bio geographical regions in which date palm growns in Niger (Figure 1). The saharan zone is generaly divided in Grand Nord and Air region while the Sahelian zone is divided in Manga and Damagaram regions. The plant material used consists of date palm leaf samples collected from Iférouane, Timia, Ingall, Bilma, Dirkou, Aguer, Aney, Achenouma, Siguidine and Djado (ten villages of Sahara in Agadez region) and from Guidimouni, Dan Kalou, Riria Loloji, Wacha, Lakiré, Jambirji, Kilboa, Anassaboul, Broumwadi and Madkwari (ten village of Sahel in Zinder and Damagaram regions). The samples were collected according to the geographical distribution of the species in order to cover the greatest possible genetic diversity, mainly in the Saharan and Sahelian zone.
Figure 1. The study area: Sahara=cercle in full line and Sahel=cercle in none full line.
Data collection: DNA was extracted from the young date palm leaves using the DNeasy Plant mini kit (Qiagen S.A., Courtaboeuf,France). DNA purification was performed with the PureLink purification kit (Invitrogen). After purification, the DNA concentrations were determined using the GeneQuant spectrometer (Amersham Pharmacia Biotech, France).
Eighteen (18) microsatellite markers Simple Sequence Repeats SSRs, Billotte et al., and one minisatellite chloroplastic marker Henderson et al., were used to collect information from 140 date palm samples of twenty (20) palm groves (one per village) in four (04)sub-regions and of two (02) main areas.
Data analysis
To compare the spatial relationship between subpopulations, we used the Principal Component Analysis (PCA) to assess the spatial genetic distribution of population units considered along spatial gradients. Principal Coordinates Analysis (PCoA) Peakall and Smouse based on the projection of genetic matrix distances constructed from the allelic frequencies based on molecular markers SSRs were performed. Genetic variation between subpopulations was evaluated using genetic differentiation index [7]. In order to analyze the relationship between date palm cultivation activities in palm groves, the geographical distribution and the genetic structure of date palm subpopulations, the Mantel test Peakall, Smouse was performed. Thus, the correlation between genetic distances and geographical distances matrices, and the one between genetic distances and the fidelity on cultural practices matrices was establish.
The matrix of genetic distances as proportion of genetic differentiation between subpopulations Fst Meirmans was determined by:
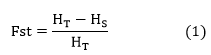
The Euclidean distance matrices from the implementation of date palm cultural practices and geographical distances were estimated by:
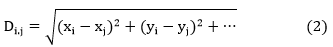
With xi and xj are respectively the values of the ith and jth individual at locus x; yi and yj are respectively the values of the ith and jth individual at locus y; HT=1-∑ki=1 Pi2 the total expected heterozygosity in Hardy Weinberg equilibrium population; HS=1-∑Pi2 the average intra-population heterozygosity; K the number of subpopulations and Pix and Piy the frequency of the ith allelic in subpopulation x or y.
The correlation between genetic distances, geographical distances and fidelity to cultural practices matrices is carried out using the simple and partial Mantel test [8].
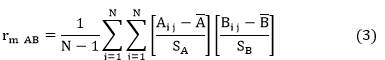
With N the number of elements of the upper or lower triangular part of the matrix, A and B are two distance matrices, A and B are respective averages of matrices A and B; SA and SB are of respective standard deviation matrices of A and B, and rm AB the Mantel correlation coefficient. The value of an association parameter or a correlation coefficient, between the two matrices is calculated from the real data, then compared to the series of pseudo-values get from random permutation. This permutation test was carried out assuming that the considered matrices A and B have the same size (n × n), the rows and columns correspond to the same objects.
Permutation design: The following steps were used:
The tests and analysis of this study are carried out in the R version 3.6.0 environment R Development Core Team.
Genetic distribution and spatial anthropic activities
The Principal Component Analysis (PCA) carried on the subpopulation according to the implementation of cultural practices allows to see that the fidelity of cultural practices shows globally two groups mainly formed based on the multiplication mode which is seems a very important factor for genetic diversity (Figure 2).
Figure 2. Distribution of subpopulations according to the fidelity of cultural practices.
The Principal Coordinates Analysis (PCoA) on the matrix of genetic distances between subpopulations shows that the first two principal components subdivide the subpopulations into two main geographical groups (Figure 3). The first group is dominating by the Sahelian date palm and the second one is dominating by the Saharan date palm (Figure 4).
But the PCoA by detailing a little the geographical origin in sub-regions (four sub-regions) including Damagaram and Manga (Sahel); the North (G_Nord) and Aïr (Sahara) shows a structure in three geographically distributed populations with a group of palm groves of Damagaram and Manga, another formed of the palm groves of the North and at the limit a third made up of Aïr palm groves (Figure 5). This grouping follows a more spatial organization [9].
Figure 3. Projection of the genetic distribution according to the main date palm areas: Sahara (Iférouane, Timia, Ingall, Bilma, Dirkou, Aguer, Aney, Achenouma, Siguidine and Djado); Sahel (Guidimouni, Dan Kalou, Riria Loloji, Wacha, Lakiré, Jambirji, Kilboa, Anassaboul, Broumwadi and Madkwari).
Figure 4. Projection of the genetic distribution according to the date palm sub-regions: Air (Iferouane, Ingall and Timia); G_Nord (Bilma, Dirkou, Aguer, Aney, Achenouma, Siguidine and Djado); Damagaram (Guidimouni, Dan Kalou, Riria Loloji, Wacha, Lakiré, Jambirji) and Manga (Kilboa, Anassaboul, Broumwadi and Madkwari).
Figure 5. Genetic structure of the different subpopulations based on the Fst matrix.
Relationship between geographical distance and genetic variation
The Mantel test shows, a positive and significant correlation between genetic and geographic distances with a correlation coefficient rm=0.280 (P-value=0.000). The genetic differentiation (Fst) increases with geographical distance. The theoretical genetic variation (0.0174) compared to the observed genetic variation (16.108) under Monte-Carlo permutation at 100000 by Mantel test (Figure 6), shows a considerable difference (P-value=0.000) with the Mantel correlation coefficient between the matrix of geographic distances and the matrix of genetic distances between the population units considered rm=0.28. The linear correlation between the matrix of genetic distances and geographic distances is not significant (P-value=0.391, [10]. This result shows that the relationship between genetic distance and geographic distance is not a causal relationship (Figure 7).
Figure 6. Differentiation of the population into genetic groups explained by the geographical distance by the Monte-Carlo test with 100,000 bootstraps.
Figure 7. The genetic distance according to the geographical distance between subpopulations. The line corresponds to the regression line calculated. Fst=Genetic differentiation index, Geodist=Geographical distance.
Relationship between cultural practices fidelity and genetic variation
The results of studies on the Fst have already shown a decrease in the rate of heterozygosity in populations from North to South-East because in general the Fst grow from North to South-East (Figure 5)[11]. In this part, the correlation between Fst values and theindex of the implementation of cultural practices within thedifferent subpopulations (palm groves) were estimate. From theMantel test, a positive and significant correlation was revealedbetween genetic distances and the index of implementation of cultural practices with a correlation coefficient rm=0.534 (P-value=0.000). The Fst increases with the index of cultural practices fidelity. The theoretical genetic variation (0.078) compared to that observed variation (6.801) under the Monte-Carlo permutation at 100000, the Mantel test (Figure 8) shows a considerable difference between them (P-value=0.000) and a significant mantel correlation coefficient (rm=0.534) between the two matrices. Figure 9 shows the linear correlation between the matrix of genetic distances Fst and the matrix of cultural practices distances is significant (P-value=0.000) [12].
Figure 8. Differentiation of the population into genetic groups explained by the implementation of cultural practices by the Monte-Carlo test with 100,000 bootstraps.
Figure 9. Representation of the genetic distance according to the implementation of cultural practices between subpopulations. The line corresponds to the regression line calculated on the data set. Fst=Index of genetic differentiation, ICP=Index of cultural practices implementation.
The date palm is of great socio-economic importance in Niger [13]. The date palm also appears essential in oasis agrosystems by creating cooler and humid local climatic conditions. Thus, it allows the cultivation of fruit trees, cereals or legumes Riou and its ecosystem services also allow the development of various forms of animal species, essential for the populations of the Saharan zone [14]. Indeed, it is therefore important to know not only the genetic structure, but also the anthropic and environmental parameters that influence its genetic structure for better conservation of this biological resource. Therefore, we are interested in this work on problems of spatial scale and we are interested in the diversity within the Phoenix dactylifera, for the effective conservation of the phoenicicole ressources in Niger (West Africa). One of the main objectives of this work is to evaluate the genetic status of date palm in Niger. Any genetically homogeneous species is threatened, so it’s important, for biodiversity conservation and for genetic diversity prediction, to understand its structuring into subpopulations and the parameters that influence it. Another objective of conservation biology is to ensure that environmental changes and human activities don’t affect the ecosystem balance, however, the study of the relationship between environment and genetic structure is essential.
The genetic structure of date palm population in Niger can be firstly explained by cultural practices and secondly by the geographical distance which separates the population units. Knowledge of this population genetic structure will contribute to the search of date palm origin, which still remains a unknow in the world. One of the genetics aims is to understand how environmental heterogeneity can influence genetic structure and diversity. According to this study, the date palm population is spatially structured into genetic groups according to the biogeographical zones of Niger. This biogeographic structuring is significantly linked to geographical distance and the exercise of certain anthropic activities related to date palm management. According to the regression coefficients, the relationship between genetic and geographic distance does not constitute a cause-and-effect relationship, so this significant test indicates that the genetic variability is structured in geographic areas, as a spatial autocorrelation. But, the relationship between genetic distribution and cultural practices could be a causal relationship [15].
Maintaining genetic variability is one of the major objectives of biodiversity conservation [16]. Knowledge of inter- and intra-population genetic variation provides essential information for the formulation of good direct management strategies for the conservation of biological resources Milligan et al. both in situ and ex situ. Given that the genetic structure of this species exhibits genetic diversity and gene exchange in the population suggest that genetic drift is currently not of great concern for this species. The discovery of pollen flow and the effects of population structure of this species, population fragmentation is associated with population size, decreases and increases the geographic separation of populations, reducing its success rate of breeding [17]. However, assessing the risk of population extinction is a central problem in conservation biology. As we saw in the introduction part, geographically isolated populations are subject to environmental variations, and can suffer of demographic and genetic problems that can increase their risk of extinction. Genetic problems are linked to the phenomena of drift leading to a loss of genetic diversity. The level of genetic diversity is often measured in natural populations by the level of individual heterozygosity and by the number of alleles observed at neutral nuclear loci. It is shown that data on genetic variability at nuclear markers can provide indications of extinction risks [18]. Moreover, these patterns of genetic variability, resulting from past and present characteristics of demographic events, can provide information on the risks of extinction that are difficult to obtain with purely ecological or demographic analyses, although this link between the level of genetic variability and extinction risk is not clearly established, some studies have shown negative correlations between heterozygosity and species extinction risk [19]. The Fst values are variable between date palm subpopulations in Niger, which suggests that the risk of extinction also varies between these population units. Therefore, from one biogeographical zone to another and according to the model of cultural practices carried out in the palm groves, we can predict the genetic diversity and the origin of the sub-population [20].
Date palm subpopulations in Niger are structured into biogeographically distributed groups. The geographical distances between the palm groves and those of implementation of certain cultural practices like, modes of multiplication, types of pollination can explain the genetic differentiation between these population units. Thus, the distance of the implementation of cultural practices influences more considerably the genetic structuring of the date palm in Niger. This study shows that the anthropic activity related to the date palm management carried out in the palm groves and the geographical distribution of the palm groves are the best predictors of the genetic distance between the population units of the date palm in Niger. Date palm genetic resources are part of the national heritage of several countries in the world, its use and conservation must therefore be the subject of explicit national policies. Hence, it’s very interesting to consider at the national, regional and international global level of conservation for genetic diversity and biological resources concerning species whose range extends over several countries such as the date palm and especially of great socio-economic and ecological importance, and also very adapted to very precarious climatic conditions.
This research was financed by Sud Expert Plantes Development Durable (SEP2D) program from the Institute of Research for Development (IRD/France) under the Project AAP2-146 “Date Palm Agro-Biodiversity in Sahel”. Thanks to farmers and village leaders who have agreed to work with us. Authors would like also to thank ONG GASSAR, Oasis Development Support Association in Niger for the facilities.
The authors declare that they have no conflict of interest.
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
[Crossref] [Google Scholar] [PubMed]
Select your language of interest to view the total content in your interested language